
Sampling around points of interest
2025-02-11
Source:vignettes/articles/guide_sample_lsm.Rmd
guide_sample_lsm.Rmd
library(landscapemetrics)
library(terra)
library(sf)
library(ggplot2)
# internal data needs to be read
landscape <- terra::rast(landscapemetrics::landscape)landscapemetrics provides several functions to sample metrics at or around sample points. On possible application for this feature could be a study in which the study organism only encounters the landscape within a local neighborhood of sample points. For most functions, sample points can be provided as a 2-column matrix(x- and y-coordinate), or sf objects (Pebesma 2018) are supported.
First, we create some example sample locations.
# create some example points
(points <- matrix(c(10, 5, 25, 15, 5, 25), ncol = 2, byrow = TRUE))
## [,1] [,2]
## [1,] 10 5
## [2,] 25 15
## [3,] 5 25
# # create some example lines
# x1 <- c(1, 5, 15, 10)
# y1 <- c(1, 5, 15, 25)
# x2 <- c(10, 25)
# y2 <- c(5, 5)
#
# sample_lines <- sf::st_multilinestring(x = list(cbind(x1, y1), cbind(x2, y2)))Extract landscape metrics at sample points
extract_lsm() returns the metrics of all patches in
which a sample point is located. However, since this only makes sense
for individual patches, it’s only possible to extract patch-level
metrics.
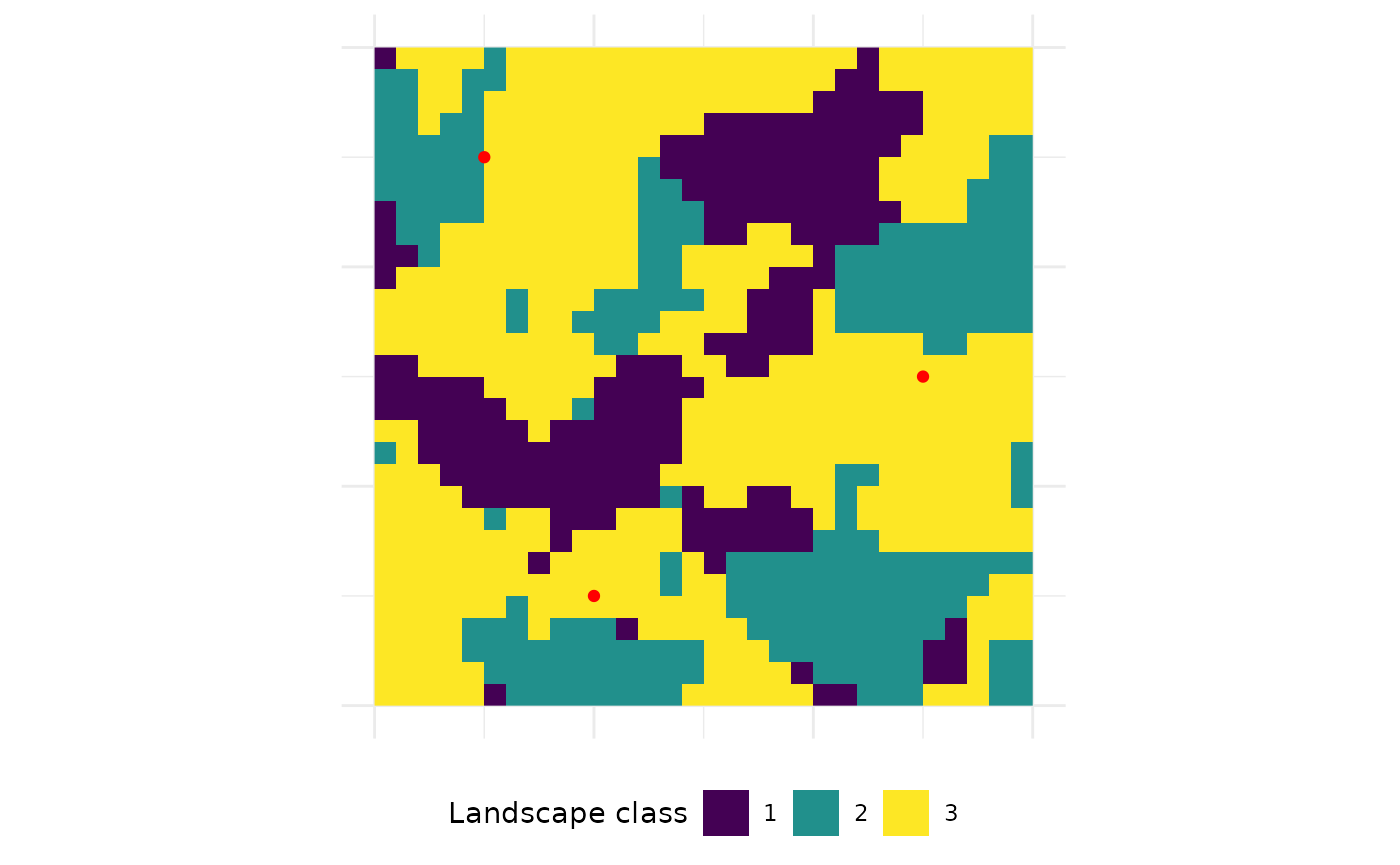
Now, it’s straightforward to extract e.g. the patch area of all
patches in which a sample point is located. Similar to all functions
calculating several landscape metrics, the selected metrics can be
specified by various arguments (see list_lsm() for more
details). The resulting tibble includes one extra column (compared to
calculate_lsm()), indicating the ID of the sample
points.
Because three sample points were provided and only the patch area was requested, the resulting tibble also has three rows - one for each sample point. The first and the second sample point are actually located in the same patch and following also the area is identical. The third sample point is located in a much smaller patch. The tibble gives also information about the patch ids and the land-cover classes in which sample points are located.
extract_lsm(landscape, y = points, what = "lsm_p_area")
## Warning: Please use 'check_landscape()' to ensure the input data is valid.
## # A tibble: 3 × 7
## layer level class id metric value extract_id
## <int> <chr> <int> <int> <chr> <dbl> <int>
## 1 1 patch 3 24 area 0.0113 1
## 2 1 patch 3 26 area 0.0148 2
## 3 1 patch 3 23 area 0.0159 3Sample landscape metrics at sample points
To sample landscape metrics within a certain buffer around
sample points, there is sample_lsm(). Now, the size of the
buffers around the sample locations must be specified. The functions
clips the landscape within the buffer (in other words sample plots) and
calculates the selected metrics.
The resulting tibble includes two extra columns. Again,
the id of the sample point is included. Furthermore, the size of the
actual sampled landscape can be different to the provided size due to
two reasons. Firstly, because clipping raster cells using a circle or a
sample plot not directly at a cell center lead to inaccuracies.
Secondly, sample plots can exceed the landscape boundary. Therefore, we
report the actual clipped sample plot area relative in relation to the
theoretical, maximum sample plot area e.g. a sample plot only half
within the landscape will have a
percentage_inside = 50.
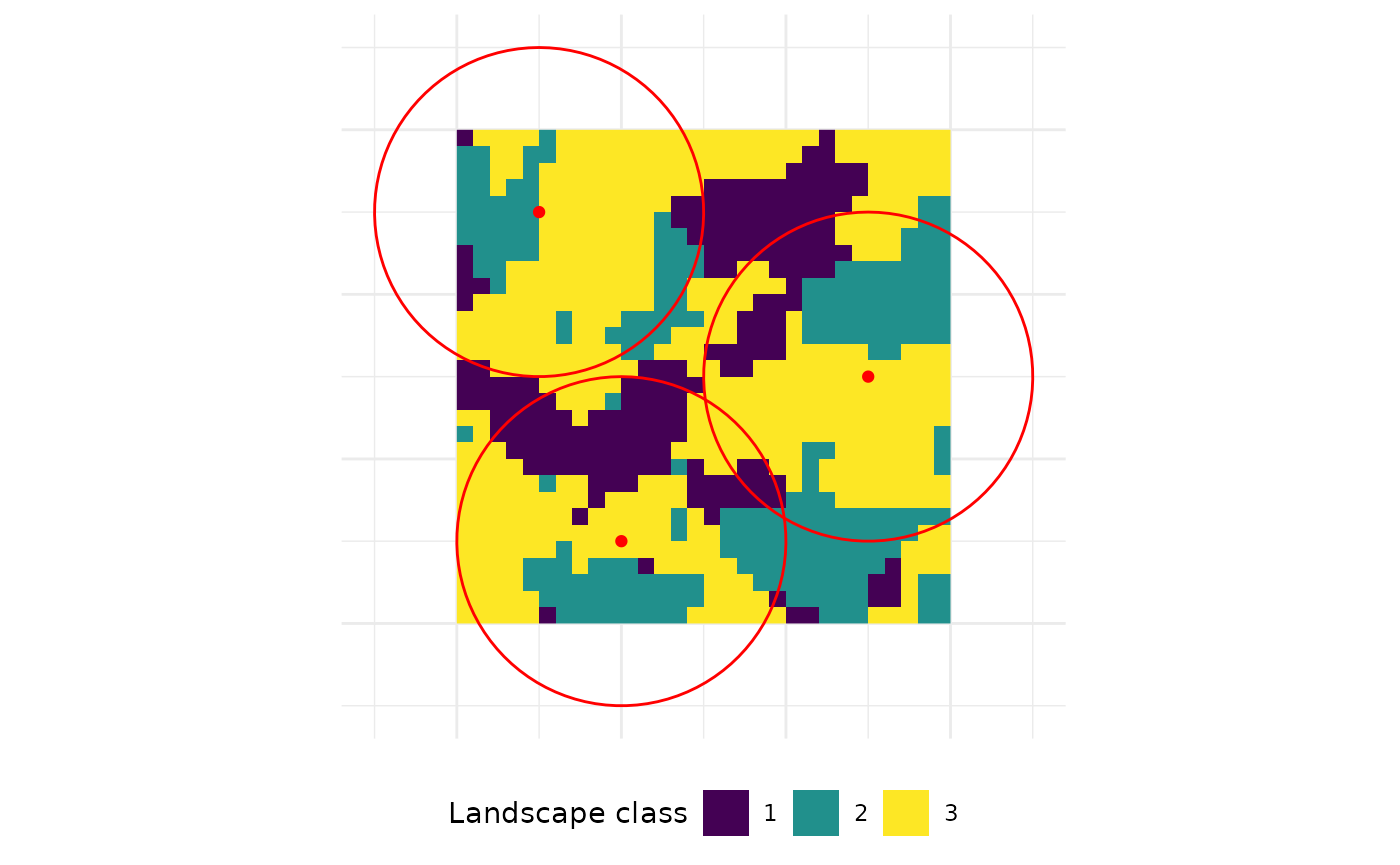
sample_lsm(landscape, y = points, size = 10,
level = "landscape", type = "diversity metric",
classes_max = 3, verbose = FALSE)
## # A tibble: 27 × 8
## layer level class id metric value plot_id percentage_inside
## <int> <chr> <int> <int> <chr> <dbl> <int> <dbl>
## 1 1 landscape NA NA msidi 0.947 1 75
## 2 1 landscape NA NA msiei 0.862 1 75
## 3 1 landscape NA NA pr 3 1 75
## 4 1 landscape NA NA prd 10000 1 75
## 5 1 landscape NA NA rpr 100 1 75
## 6 1 landscape NA NA shdi 1.02 1 75
## 7 1 landscape NA NA shei 0.928 1 75
## 8 1 landscape NA NA sidi 0.612 1 75
## 9 1 landscape NA NA siei 0.918 1 75
## 10 1 landscape NA NA msidi 0.969 2 75
## # ℹ 17 more rowsReferences
Pebesma, E., 2018. Simple Features for R: Standardized Support for Spatial Vector Data. The R Journal, https://journal.r-project.org/archive/2018/RJ-2018-009/
Roger S. Bivand, Edzer Pebesma, Virgilio Gomez-Rubio, 2013. Applied spatial data analysis with R, Second edition. Springer, NY. http://www.asdar-book.org/